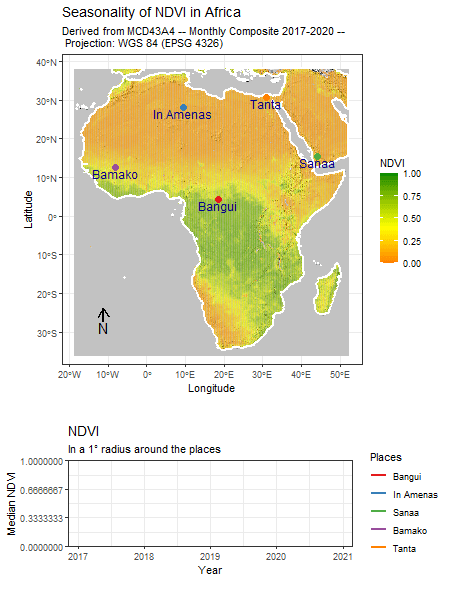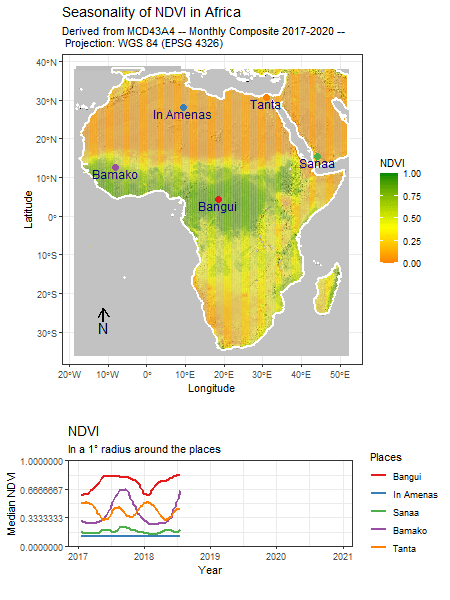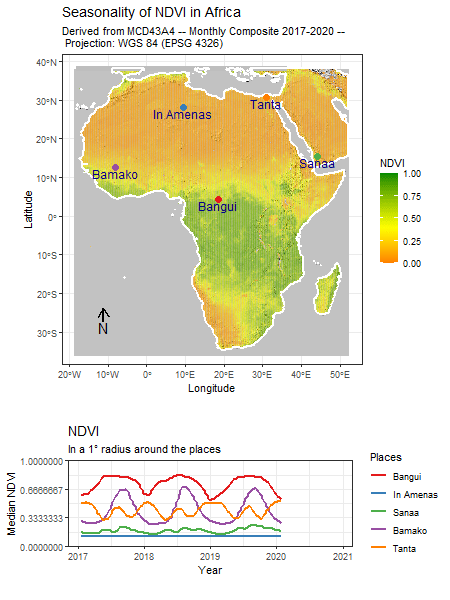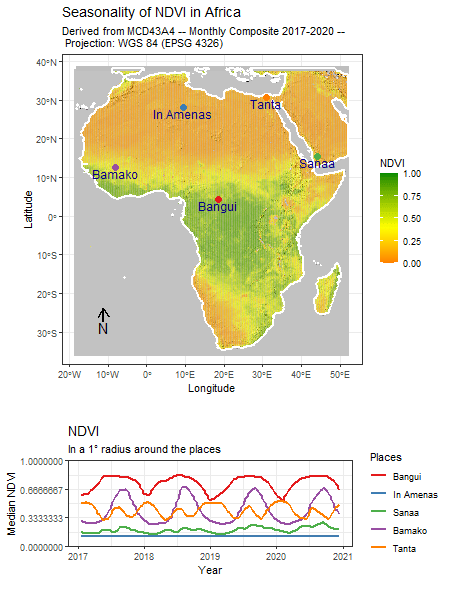
Tag: R
fieldRS updated (v0.2.1)
fieldRS 0.2.1 is on CRAN! See the package news for details. Highlights:extractFields() can...
Read MorersMove Updated! (v0.2.7)
by EORhub | Dec 12, 2018 | Biodiversity, Conservation, News, R, Software | 0 |
New version of rsMove is available for download. Check the news of the package for details. The...
Read MoreRemote Sensing and GIS for Ecologists book now available
by Martin Wegmann | Feb 3, 2016 | Biodiversity, Conservation, News, publication, Software | 0 |
Our book “Remote Sensing and GIS for Ecologists – Using Open Source software” is...
Read Moreunsupervised classification with R
Here we see three simple ways to perform an unsupervised classification on a raster dataset in R. I will show these approaches, but first we need to load the relevant packages and the actual data. You could use the Landsat data...
Read Moreupdated RStoolbox version
by EORhub | Jan 29, 2016 | Biodiversity, Conservation, News, R, Software | 0 |
The RStoolbox R package has been updated after some testing in courses and by colleagues. Please update your package using update.packages() or install the RStoolbox again. New functions are: new function `validateMap()` for...
Read MoreUpdate: R spatial handling cheatsheet
by Martin Wegmann | Oct 5, 2015 | Biodiversity, Conservation, News, R, Software | 0 |
We updated our cheatsheet on spatial data handling in R. Beside some minor changes of common raster commands we also added now some of the new RStoolbox commands within a Remote Sensing operations section such as the command for...
Read MoreRemote Sensing and GIS for Ecologists – book submitted
by Martin Wegmann | Apr 23, 2015 | Biodiversity, Conservation, News, publication, training | 0 |
We finally submitted our book “Remote Sensing and GIS for Ecologists” to the publisher. It took longer than anticipated and we learned a lot. Looking forward to the printed version. More details how to order...
Read More
news blog by
the Earth Observation Research cluster
EORC
at the University of Würzburg
Institute of Geography and Geology
John Skilton Str 4a
97074 Würzburg
recent posts
-
 New PhD student Tamilwai KolowaJul 5, 2025 | staff
New PhD student Tamilwai KolowaJul 5, 2025 | staff -
 Course on Object-based image analysisJul 4, 2025 | News
Course on Object-based image analysisJul 4, 2025 | News -

-

-